Even the smallest creatures have big stories to tell
New research uncover a conservative route to genome compaction in the microscopic segmented worm, Dimorphilus gyrociliatus, shown to possess one of the smallest free-living animal genomes with only 73.8 million base pairs.
The miniaturized genome of Dimorphilus gyrociliatus retains traits classically associated with larger and slower-evolving genomes, such as an ordered, intact Hox cluster, a generally conserved developmental toolkit, and canonical features of genome regulation. This shows an alternative pathway to genome compaction not triggering drastic changes in the genomic structure and function as reported for Nematoda (roundworms), Tardigrada (waterbears) and Urochordata (sea squirts and larvaceans), hereby questioning the assumed causal link between fast evolving genomic traits and genome compaction.
Katrine Worsaae, Department of Biology, University of Copenhagen, is in her research group conducting experiments with microscopic animal lineages for deciphering the development and evolution of metazoan body plans.
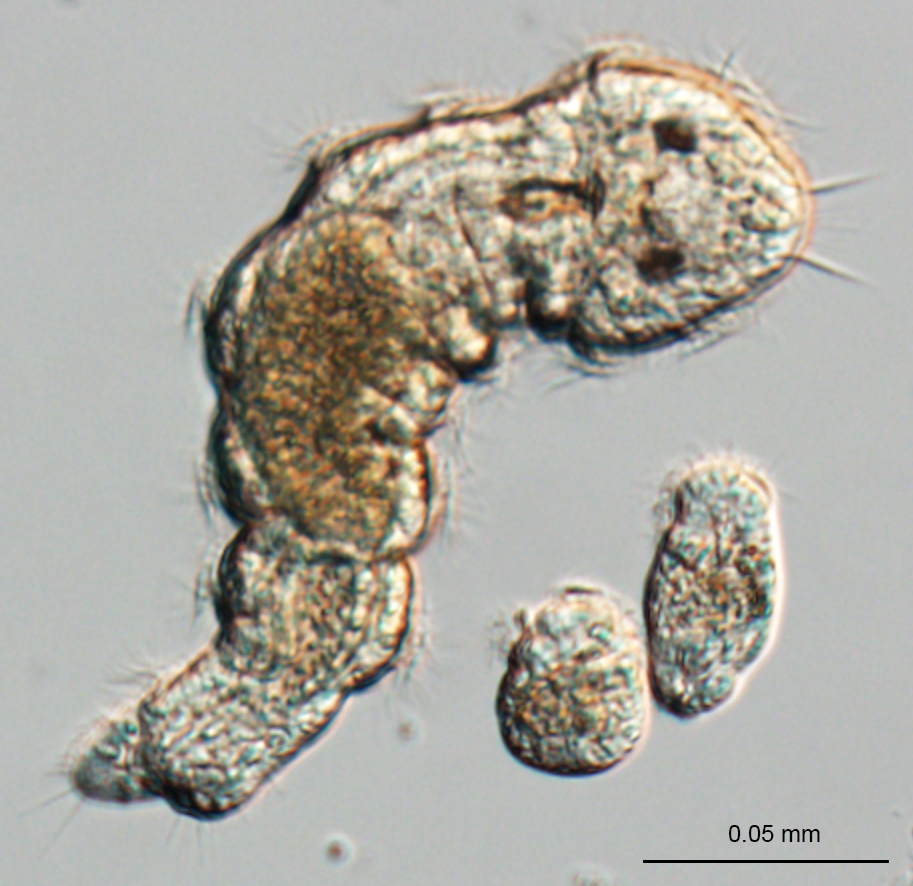
- “We unknowingly began to approach this question when I in 2012 visited Andreas Hejnol’s lab in Bergen (Norway), where Chema Martín-Duran was doing his postdoctoral research at that time. I brought with me a few 50 mL falcon tubes speckled with hundreds of crawling Dimorphilus gyrociliatus. These simple marine worms had long been my lab companions, the 1 mm long females living happily from spinach and with a fast and peculiar life cycle, incorporating some of the smallest males in the animal kingdom – only 0.05 mm long. I wanted to establish gene expression techniques together with Chema and Andreas, who’s lab already was renowned for their molecular studies on microscopic non-model organisms. We started generating transcriptomic and genomic data alongside mapping their development and nervous system, involving several students from our two laboratories”, explains Katrine Worsaae.
Animal genomes can vary nearly four orders of magnitude in size, but the scientists still understand little of why and how this happens.
- “So, when our analysis of that sequencing dataset uncovered one of the smallest animal genomes ever sequenced we decided to establish a larger international team of organismal and molecular experts, to test whether this compaction resembled that found in some other fast-evolving animal lineages”, continues Katrine.
Combining classical organismal and modern molecular approaches, they could demonstrate that genome compaction in the tiny Dimorphilus gyrociliatus largely follows a different route to that of other known animals with small genomes. Moreover, they found that changes in the MYC pathway, a core regulator of cell size and cell number in animals, could be at the core of the peculiar biology of this miniature annelid worm. Altogether, the study demonstrated a conservative route to genome compaction, highlighting once more the importance of exploring animal diversity to gain insights into some of the more puzzling questions in Biology. Because sometimes, even the smallest creatures have big stories to tell.
The research is funded by Villum Foundation and the Independent Research Fund Denmark.
Kontakt
Associate professor Katrine Worsaae
Marinbiological Section, Department of Biology
University of Copenhagen
E-mail: kworsaae@bio.ku.dk
Phone: + 45 3532 1987
Helle Blæsild
PR & Communication
Department of Biology, University of Copenhagen
E-mail: helleb@bio.ku.dk
Phone: +45 2875 22076
