BioMatrix

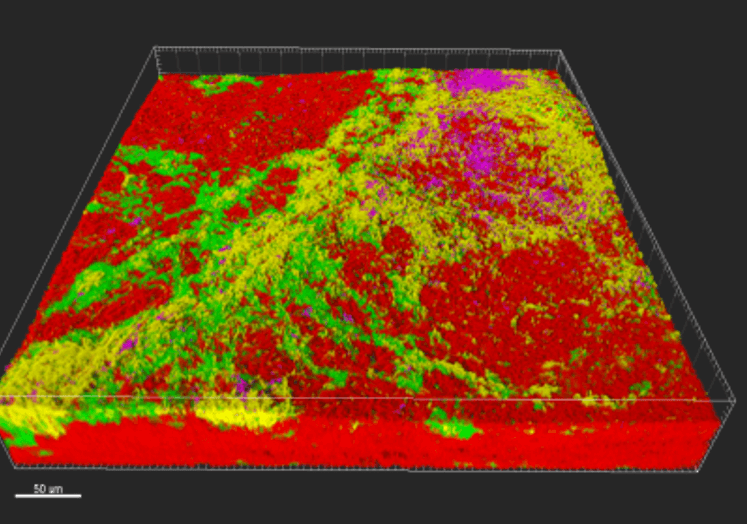
The four BioMatrix model bacterial species forming biofilm on a plant root.
Bacteria in nature live in biofilms. These biofilms are found everywhere; on plant roots, on decomposing organic material, in aquatic sediments and much more. Biofilms are useful in biotechnological applications, but they are also problematic, as they form at unwanted sites in industrial settings and can lead to chronic infections in humans.
Biofilms are assemblages of bacteria embedded in a matrix, produced by the bacteria themselves. When different types of bacteria live in proximity in a biofilm, we often identify features of this microbial community, which neither of the bacteria possess individually; examples include enhanced tolerance towards stress and inhibitors, and new metabolic capabilities. The interactions and mechanisms causing such “community properties” are largely unknown.
In BioMatrix, we will examine the matrix of biofilms comprised of multiple types of bacteria. We aim at identifying matrix components that directly impact functionality and/or tolerance of biofilms in nature. New experimental models will be designed to better mimic the real life of biofilm bacteria in nature.
BioMatrix will uncover fundamental mechanisms shaping bacterial communities and thereby close a primary knowledge gap in microbial ecology. Moreover, specific targets, facilitating efficient biofilm manipulation, are provided. This is useful in biotechnology for enhanced exploitation of bacterial community activity and in the development of novel and improved chronic infection treatment strategies.


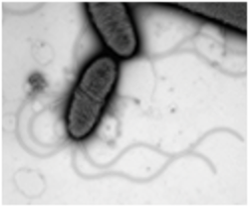
- Sadiq, F.A., Yang, N., Goeteyn, J. et al. Microbial Interactions Shape Spatial Organisation and Transcriptional Responses in a Model Mixed-Species Biofilm. Microb Ecol 89, 65 (2026). https://doi.org/10.1007/s00248-026-02701-w
- Ronin, D., Hansen, M.F., Flaig, M.L., Dueholm, M.K.D., Hostrup Daugberg, A.O., Nesme, J., Kot, W. & Burmølle, M. Unfolding the collective functional potential of a synergistic multispecies community through genotypic and phenotypic analyses. Biofilm 10, 100290 (2025). https://doi.org/10.1016/j.bioflm.2025.100290
- Amador, C. I., Røder, H. L., Herschend, J., Neu, T. R. & Burmølle, M. Decoding the impact of interspecies interactions on biofilm matrix components. Biofilm 9, 100271 (2025). https://doi.org/10.1016/j.bioflm.2025.100271
- Khani, M., Kiesewalter, H. T., Hansen, M. F., Ronin, D., Nesme, J., Rasekh, B., Nosrati, M. & Burmølle, M. Enzymatic exposure impacts microbial diversity and reduces biovolume of reverse osmosis membrane‑associated biofilms. Chemical Engineering Journal 507, 160186 (2025). https://doi.org/10.1016/j.cej.2025.160186
- Ronin, D., Hansen, M.F. & Burmølle, M. Co‑cultivation rescues suicidal Paenibacillus amylolyticus swarms. The ISME Journal 19(1), wraf225 (2025). https://doi.org/10.1093/ismejo/wraf225
- Hansen, M.F., Ronin, D., Kiesewalter, H.T., Amador, C.I. & Burmølle, M. The Role of Matrix Components in Multispecies Coexistence in Biofilms. In Biofilm Matrix (Springer Series on Biofilms, Vol. 13), pp. 187–214 (Springer Nature, 2024). https://doi.org/10.1007/978-3-031-70476-5_6
- Sadiq, F.A., De Reu, K., Yang, N., Burmølle, M. & Heyndrickx, M. Interspecies interactions in dairy biofilms drive community structure and response against cleaning and disinfection. Biofilm 7, 100195 (2024). https://doi.org/10.1016/j.bioflm.2024.100195
- Yang, N., Røder, H. L., Wicaksono, W. A., Wassermann, B., Russel, J., Li, X., Nesme, J., Berg, G., Sørensen, S. J. & Burmølle, M. Interspecific interactions facilitate keystone species in a multispecies biofilm that promotes plant growth. The ISME Journal 18(1), wrae012 (2024). https://doi.org/10.1093/ismejo/wrae012
- Sadiq, F.A., De Reu, K., Steenackers, H., Van de Walle, A., Burmølle, M. & Heyndrickx, M. Dynamic social interactions and keystone species shape the diversity and stability of mixed‑species biofilms – an example from dairy isolates. ISME Commun. 3, 118 (2023). https://doi.org/10.1038/s43705-023-00328-3
- Khani, M., Hansen, M.F., Knøchel, S., Rasekh, B., Ghasemipanah, K., Zamir, S.M., Nosrati, M. & Burmølle, M. Antifouling potential of enzymes applied to reverse osmosis membranes. Biofilm 5, 100119 (2023). https://doi.org/10.1016/j.bioflm.2023.100119
- Deyong Zhu
- Nan Yang
- Lutian Wang
- Keiko T Kiesewalter
- Dana Ronin
- Joana Azevedo
- Gustav Olsen
- Mojtaba Khani
- Johannes Højlund Olsen
- Faizan Sadiq (visiting Marie Curie Postdoc)
Contact

Professor Mette Burmølle
Section of Microbiology
Universitetsparken 15,
Building 1, 1. floor
DK-2100 Copenhagen
Mobile: +45 4022 0069
Mail: burmolle@bio.ku.dk
Funded by:

BioMatrix is a project ERC Horizon 2020 funded consolidator grant to PI Mette Burmølle.
 The BioMatrix Project
The BioMatrix Project
Project team
| Name | Title | Phone | |
|---|---|---|---|
| Cristina Isabel Amador Hierro | Associate Professor | +4535334850 | |
| Mads Frederik Hansen | Postdoc | +4535335180 | |
| Maximilian Lukas Flaig | PhD Fellow | ||
| Mette Burmølle | Professor | +4535330119 | |
| Siwei Liang | PhD Student |
Students
| Name | Title |
|---|---|
| Anna Maia Berg Gudiksen | MSc student |
| Freja Sandager Carlsson | MSc student |
| Maria Palomeque Sanchez | MSc student |
