Plant Microbiome Lab – Flemming Ekelunds Group
we are interested in protozoa, an often-neglected group of eukaryotic microorganisms that play an essential role for plant nutrition. At present, our main focus is how diversity of soil bacteria, and especially the trophic interactions between soil bacteria and the protozoa that feed on them can be optimised to support plant growth.
We work with state of the art RNA-based techniques, which allow concomitant mapping of the entire microbiome and the complementary plant transcriptome. It is our vision that the more complete picture, we provide of plant- and microbial traits, and their interaction, will help facilitate the green transition.
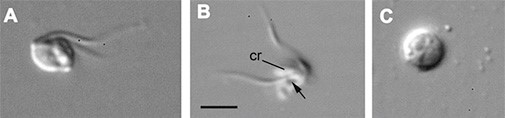
The protozoan (flagellate) Otto terricolus (named after my father Otto), which I discovered in a soil sample from Foulum. A and B show swimming flagellates, C shows a resting stage, a cyst. Otto terricolus is common in soil, but is very small (the scale bar is 3 µm = 3/1000 mm); probably the reason why it was only discovered a few years ago. The flagellate feeds on bacteria, which stimulates nutrient turnover and hence plant growth.
 Barley plants. In the project Plant-microbiome interactions in modern and ancient barley, we aim to optimise plant microbiomes, with special focus on bacterial-protozoan interactions, to support plant growth. We hypothesise that modern plants have lost many of the genes and pathways that provide them with the ability to interact with the microbiome. Therefore, we focus on ancient varieties. We aim at producing systems with optimal plant growth with minimal nutrient input to facilitate the green transition.
Barley plants. In the project Plant-microbiome interactions in modern and ancient barley, we aim to optimise plant microbiomes, with special focus on bacterial-protozoan interactions, to support plant growth. We hypothesise that modern plants have lost many of the genes and pathways that provide them with the ability to interact with the microbiome. Therefore, we focus on ancient varieties. We aim at producing systems with optimal plant growth with minimal nutrient input to facilitate the green transition.
Plant-microbiome interactions in modern and ancient barley
Modern plant breeding has yielded crop genotypes that need precise anthropogenic chemical control, thereby largely neglecting that the rhizosphere microbiome can protect against diseases and promote plant nutrient availability and uptake.
In the project, we investigate the extent of the plant-microbiome interactions between 6 varieties of barley (3 modern & 3 ancient) and 9 different microbiomes under 3 different nutrient regimes. We investigate to which extent modern varieties have lost the ability to interact with the microbiome with focus on plant/bacteria/eukaryotic micro-grazer interactions.
We sequence of the root/leaf transcriptome to reveal genes and pathways involved in these interactions. In parallel, we sequence the microbiome and perform multielement analyses to capture the dynamics of the interactions.
Our results will be a significant contribution to the understanding of the role of the microbiome in modern plant production. The project is housed at Department of Biology (KU) and has participants from PLEN (KU), Department of Agroecology (AU) and Globe Institute.
Protective Microbiomes on Plant Pathogenic Nematodes
In the project, we characterize microbiome components, including bacteria, fungi and protists, adhering to root-knot nematodes and investigate the microbiome’s significance for nematode survival, resistance to nemato-pathogens, and plant infectivity.
Further, we will identify microbiome functional genes involved in nematode protection and inhibition, respectively. The project is housed at Department of Agroecology (AU) and has participants from Department of Biology (KU) and PLEN (KU).
Aggregate
Soil provides multiple ecosystem services to humans of course as being the fundamental basis for agricultural food production but also by storing carbon and providing clean drinking water.
Soil aggregation correlates with SOM content, aeration and soil biodiversity. A healthy and fertile soil is well aggregated. Soil aggregation is a process where soil minerals and organic matter is enmeshed into a biological web including fungal hyphae and a matrix exuded by microbes.
In Aggregate we wish to determine which soil biota are most important for the soil aggregation process across a range of different Danish soils with focus on fungal activity and hyphae. We also want to pinpoint potential troubles for the aggregation process - in specific of pesticides. The two together will provide recommendations towards managing our agricultural soil resources in a sustainable way.
Other current research activities include
1) Bryophyte ecology and diversity. How do bryophytes interact with the ecosystem, including the microorganisms, and which factors determine bryophyte diversity?
2) Habitat ecology. How do particular animals and vascular plants affect and shape their habitats and vice versa, how do habitat conditions influence which organisms can inhabit them?
To view publications, please visit Flemming Ekelund’s profile on the KU Research Portal.
Current funding supports three research projects:
- FNU: Plant-microbiome interactions in modern and ancient barley
- FTP: Protective Microbiomes on Plant Pathogenic Nematodes
- NNF: Biological aggregate formation towards a healthy soil (AGGREGATE).
Contact Flemming if you are interested in BSc or MSc projects on Plant and/or microbiome interactions, or if you are interested in other of the topics above. If you have another idea, which may fit in here, you are welcome to contact us, then we can discuss whether it is suitable as a project. We also have a list of specific project suggestions. Check these out on UCPH Projects and Jobs - search for Flemming Ekelund.
Group members
| Name | Title | Phone | |
|---|---|---|---|
| Flemming Ekelund | Associate Professor | +4551827041 | |
| Jesper Liengaard Johansen | Assistant Professor | +4535332317 | |
| Nikolaj Lunding Kindtler | +4535330608 | ||
| Olivera Topalovic | Postdoc | ||
| Sanea Sheikh | Postdoc |
