Jakob R. Winther
My scientific work is focused on basic aspects of protein chemistry and using molecular biology and molecular genetics to understand structure-function relationships in proteins. We are mainly interested in protein engineering and design. We have developed genetic and other high-throughput screening methods as well as improved methods for using deep mutational scanning data for protein and enzyme optimization (Johansson et al, 2020, Norrild et al, 2022).
Recently, we have taken a special interest in luciferases. These are enzymes that turn over substrates (luciferins) to generate light. Luciferases have a wide distribution in nature, and are particularly prevalent on marine environments where in the order of half of all species are luminescent in one way or another. How luminescence is involved in the ecological interplay with friend and foe is mostly unknown. What is also interesting is that the basic biochemistry of light emission is mostly unknown. We are presently focusing on the luciferase from the copepod Gaussia princeps. This luciferase displays the strongest luminescence of any luciferase known, however, does so in a very short time window. We have shown that this is due to its inactivation taking place during catalysis by which it self-modifies and rapidly looses activity (Dijkema et al, 2021).
I have significant experience with teaching and student supervision and always have projects on hand. If you are interested in studying or working in my group read more about projects here and read more about joining the lab as a student on the Section for Biomolecular Sciences student page.
My scientific work is focused on basic aspects of protein chemistry and using molecular biology and molecular genetics to understand structure-function relationships in proteins.
All the projects described below are carried out in close collaboration with Martin Willemoes, Kresten Lindorff-Larsen and Kaare Teilum.
We currently work on four different projects:
- Protein design
- Development and application of strategies for selection and screening of functional proteins and enzymes
- Luciferase function and biochemistry
Protein design
In nature, proteins are able to take on a huge range of functions both as structural entities and as enzymes, the latter being the most versatile and specific chemical catalysts known. Since the 1960s it has been understood that the protein structure is solely defined by its amino acid sequence. Following this realization, a huge effort has been made to predict protein structure from sequence, however, only recently are the daunting theoretical and computational challenges relating to this problem beginning to yield.
Contemporary tools of chemistry and molecular biology allow protein sequences to be routinely generated from synthetic DNA and recent advances in the computational methods have made it possible in a few cases to design completely new proteins with specific and novel structures.
Development and application of strategies for selection and screening of functional proteins and enzymes
To be able to efficiently identify folded sequences we are developing a model system for genetic screening and selection for protein fold independent on enzymatic activity of the design target protein. In this system the folding of the target protein is linked to an enzymatic activity which is genetically selectable.
We hope that such genetic systems will enable us to select for improved designs based on the survival of E.coli that carry more efficiently folding and stable designed proteins. Such systems should also be applicable for the improvement of thermal stability of marginally stable natural proteins.
Luciferase structure function and biochemistry
Bioluminescence is a tantalizing phenomenon occurring in all kingdoms of life – from fireflies to bacteria, algae, fungi, worms and fish. The phenomenon is due to chemical reactions catalyzed by luciferase enzymes which have found widespread applications as tools in many biochemical and biomedical techniques. Surprisingly, the luminescence dynamics of luciferases have over the last 20-30 years been classified as “glow” or “flash” without clearly defining these terms mechanistically. Whereas “glow” mechanism could be explained in terms of conventional enzyme kinetics, we have recently discovered that “flash” kinetics of one of the most luminescent and widely used luciferases, from the copepod Gaussia princeps, can be explained by rapid substrate-dependent self-destruction. We are currently investigating the mechanism of this inactivation and as well a structure-function relationships in this highly abnormal enzyme.
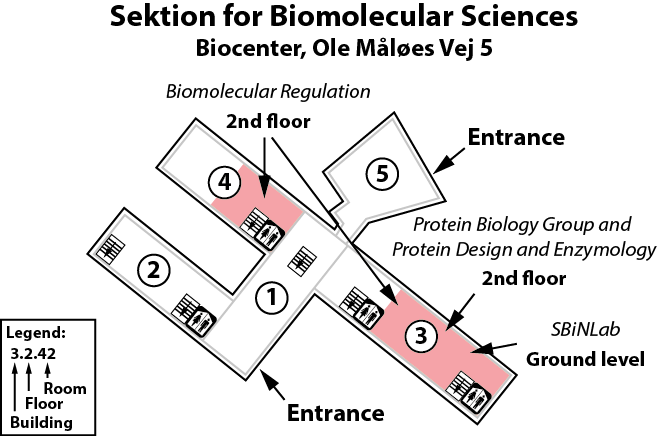
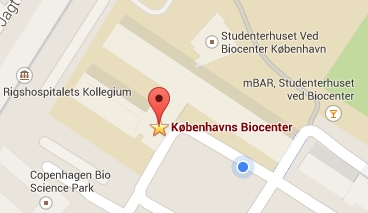
The Winther lab is located on the 2nd floor, building 3 of the Copenhagen Biocenter (map), close to the centre of Copenhagen. It takes less than 1 hour to travel from Copenhagen's airport to our lab, by train or subway (from the airport to Nørreport station, a 20 minute journey, with trains every 10 minutes) and bus (bus 150S from Nørreport to the stop called "Fredrik Bajers Plads" in the intersection of Tagensvej and Nørre Allé, a 10 minute journey, with buses every 6-8 minutes). From there it is a few minutes walk.
Contact

Professor Jakob R. Winther
Section for Biomolecular Sciences
Kaj Ulrik Linderstrøm-Lang Centre for Protein Science
Ole Maaløes Vej 5, room 3-2-31
DK-2200 Copenhagen N
Email: jrwinther_at_bio.ku.dk
Phone: +45 3532 1500
KU profile: Link