Welcome to the Skriver Group
Welcome to our website! Here, you’ll find information about ongoing research, the group, positions and how to contact us.
I have supervised many students, and I am always interested in discussing new projects with potential students or coworkers. Read more about how to join my lab on the Section for Biomolecular Sciences homepage.
The main focus of our research is gene regulatory networks and the importance of protein structure and function to the networks. More specifically, we study gene regulation mediated by specific transcription factors and the function of protein intrinsic disorder in the networks. In our studies, we integrate a number of methods from protein chemistry, biochemistry, and molecular biology. We have a strong collaboration with groups within and outside the Department of Biology both in academia and industry.
Research Activities in biochemistry
Living cells depend on regulated gene expression, which in turn depends on recognition of specific promoter sequences by gene specific transcription factors. Gene regulatory networks describe how transcription factors associate with downstream target genes. These networks can be expanded to describe the complex links with upstream regulators and between numerous proteins. The understanding of how living organisms function depends on unraveling of these biological networks. The overall objective of the research in our group is to increase the understanding of specific networks. In particular, we investigate how cells react to stress exposure at the molecular level, and how molecular structure is important to these reactions. Currently, we investigate the following:
- How do stress-associated transcription factors target specific genes, and what are the target genes of importance to stress responses?
- How do stress-associated members of the NAC transcription factor family interact with other proteins, and how do these interactions lead to agonistic and antagonistic cooperation?
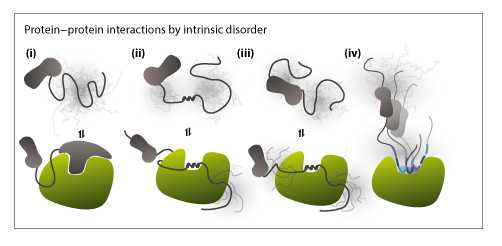
- How does protein intrinsic disorder contribute to regulatory interactions of stress-associated transcription factors, and how can knowledge about protein intrinsic disorder obtained in vitro be translated to the in vivo organismal level?
Methods used
Quantitative real-time PCR, applied bioinformatics, chromatin immunoprecipitation, protein binding microarrays, protein production and purification, biochemical assays, fluorescence and circular dicroism spectroscopy, isothermal titration calorimetry and transgenic complementation.
Research Activities in didactics
We investigate different problems of conceptual understanding that students have in biochemistry, and we experiment with new teaching styles to improve this understanding. This includes development of virtual exercises - not to eliminate hands-on lab experiences in biochemistry, but either as preparation for lab work or as a supplement to theoretical exposition and exercises.
Collaborators
Birthe Kragelund, Dept. Biology: Structure determination
Per Gregersen, University of Aarhus: Crop NAC transcription factors
Fred de Masi, Technical University of Denmark: Protein binding microarrays
Genevieve Thon, Dept. Biology: Yeast protein structure function
Klaas Vandepoele, Gent University: Bioinformatics
Leila Lo Leggio, Dept. Chemistry: NAC transcription factors
Martin Willemoes, Dept. Biology: Calorimetry
Michael May, Dept. Science Education: Didactics
Tamo Fukamizo, Kinki University: Chitinases
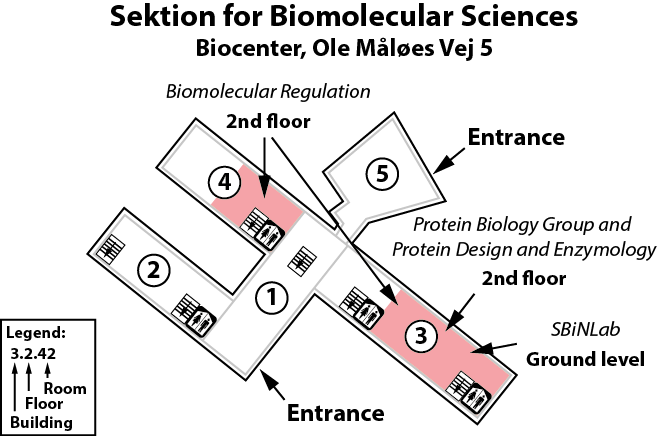
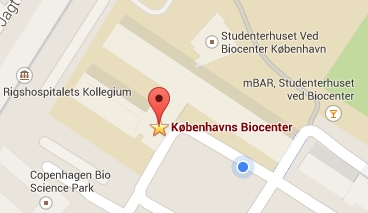
Our lab is located on the 2nd floor, building 3 of the Copenhagen Biocenter (map), close to the centre of Copenhagen. It takes less than 1 hour to travel from Copenhagen's airport to our lab, by train or subway (from the airport to Nørreport station, a 20 minute journey, with trains every 10 minutes) and bus (bus 150S from Nørreport to the stop called "Fredrik Bajers Plads" in the intersection of Tagensvej and Nørre Allé, a 10 minute journey, with buses every 6-8 minutes). From there it is a few minutes walk.
Contact

Professor Karen Skriver
Section for Biomolecular Sciences
Kaj Ulrik Linderstrøm-Lang Centre for Protein Science
Ole Maaløes Vej 5, room 3-2-43
DK-2200 Copenhagen N
Email: kskriver_at_bio.ku.dk
Phone: +45 3532 1712
KU profile: Link